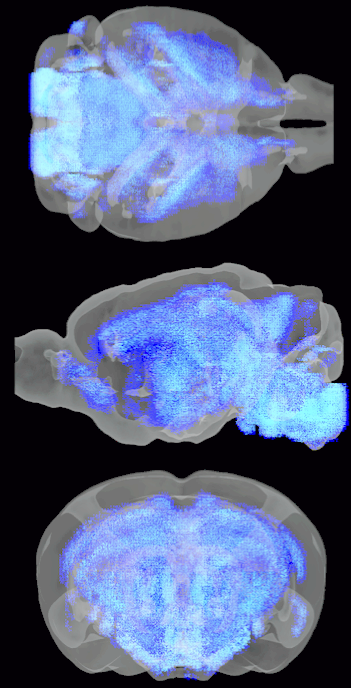
An estimated distribution of oligodendrocytes across the mouse brain.
Cell Type Mapping
Collaborators and I are working with several pre-clustered scRNAseq datasets to map the density of each cell-type cluster, using the Allen Institute Mouse Brain ISH Atlas and an extended version of the Human Brain Microarray Atlas as our spatial transcriptomes. We are currently fine tuning a procedure that determines the subset of genes, which is selected using our algorithm, MRx3, that optimally reclusters cells into their types according to their scRNAseq data and then applies non-negative linear regression to this subset of genes in the ISH & Microarray atlases to deconvolve the densities of different types of neuronal and non-neuronal cells across all voxels in both the mouse and human brain. This project is currently undergoing a major revision, but a pre-revision version of the manuscript can be found on BioRxiv.
This image was constructed using Brainframe, as were all other 3D brain renderings on this page.
Brainframe
A collaborator and I have been working on a MATLAB package that can render any arbitrary 3D brain volume from any species. All of the 3D brain renderings on this page were produced with code from this package, which is named Brainframe. We have a GitHub page for the current package, which has been tested with mouse brains using the Allen Institute CCF Volumetric Atlas and with human brains using the Desikan Atlas. We are going to test the package in rats and zebrafish soon.
Whole-brain seeding followed by connectome spread (right column) can recreate tau pathology data (left column) from a Psen1 x APPswe x P301S mouse model of disease.
Predicting Tauopathy & Synucleinopathy Using an Extended Network Diffusion Model
There is a plethora of prior work, including some of my prior work, demonstrating that network diffusion and transmission models of pathology spread can accurately recreate the spatiotemporal development of degenerative diseases. Collaborators and I are currently working on a network model that incorporates data about the spatial expression patterns of various genes and distributions of different types of cells across both mouse and human brains alongside whole brain connectomes. We are hoping to help determine to what extent pathology spread is determined by the connectome versus the degree to which some parts of the brain might be selectively vulnerable or invulnerable to pathology. Mouse data underlying the visualization in the left column pictured above from: Hurtado, DE et al. 2010. A-beta accelerates the spatiotemporal progression of tau pathology and augments tau amyloidosis in an Alzheimer mouse model. The American Journal of Pathology, Volume 177(4), pp. 1977-1988.
Cell bodies selected for calcium imaging in a zebrafish brain.
Zebrafish Functional Connectomes
I’m helping a collaborator reconstruct calcium imaging timeseries, and the functional connectome that optimally explains the reconstructed timeseries, using Dynamic Mode Decomposition (DMD). We are in the process of drafting a manuscript demonstrating the accuracy of DMD recreated calcium imaging timeseries in head-fixed zebrafish. Additionally, we are working to determine how the functional connectome changes between largely motionless versus tail whipping behavioral states.